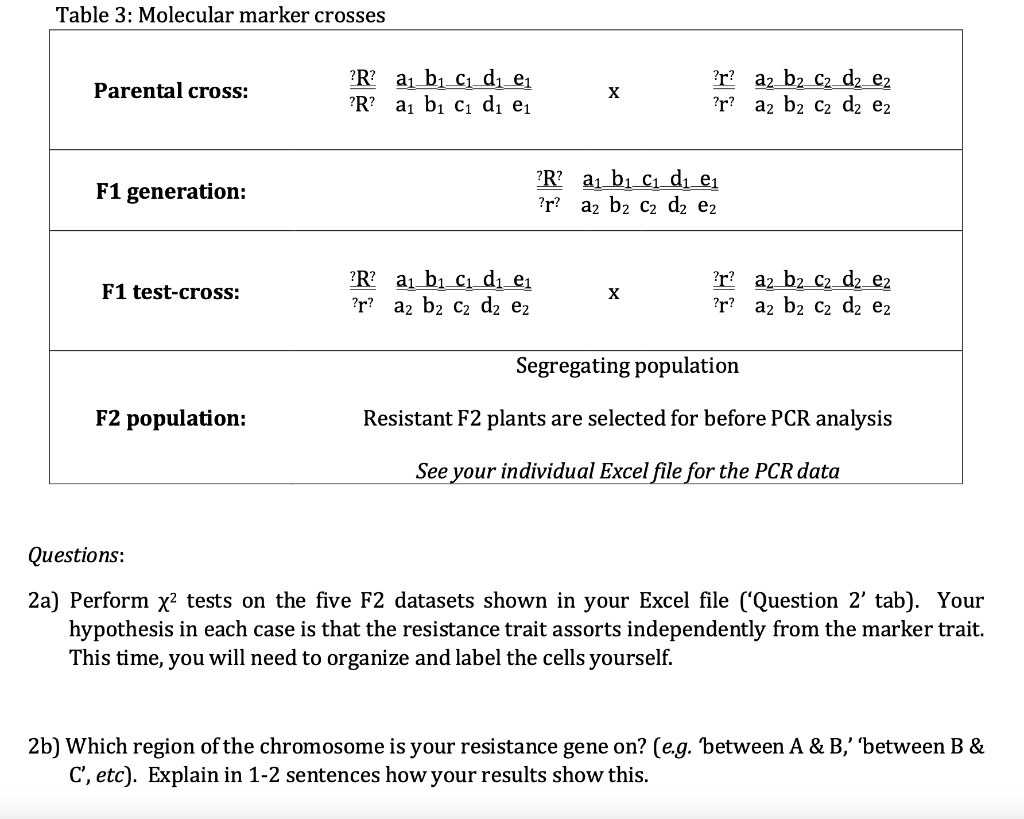
Biparental QTL mapping. a Single marker analysis. The red horizontal... | Download Scientific Diagram

Effect of Trait Heritability, Training Population Size and Marker Density on Genomic Prediction Accuracy Estimation in 22 bi-par

PDF) Bridging the genotyping gap: using genotyping by sequencing (GBS) to add high-density SNP markers and new value to traditional bi-parental mapping and breeding populations | Susan McCouch - Academia.edu

Multi-parent populations in crops: a toolbox integrating genomics and genetic mapping with breeding | Heredity
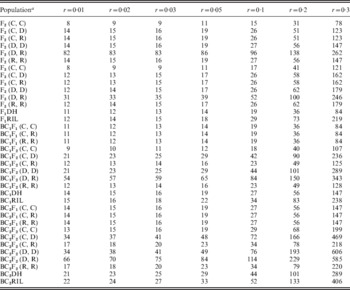
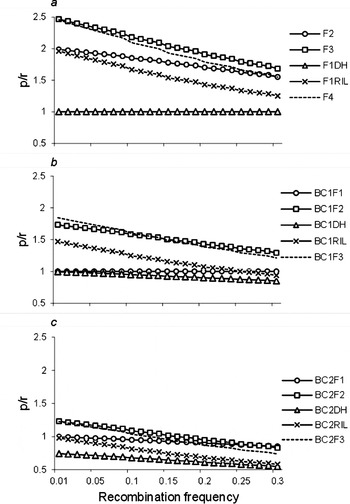
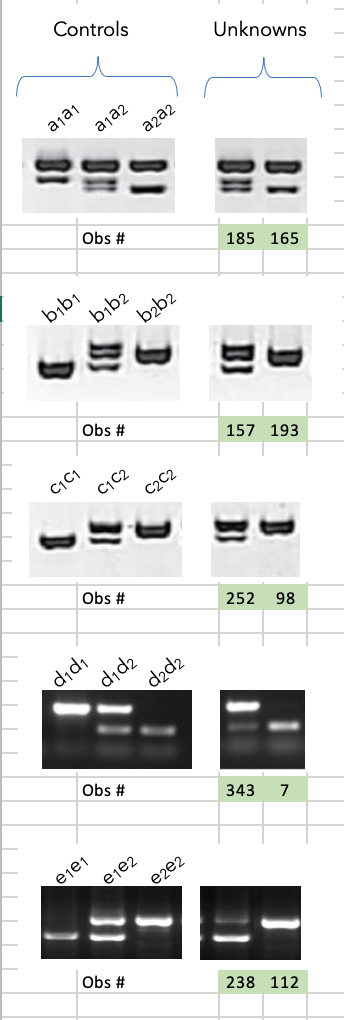
Estimation of recombination frequency in bi-parental genetic populations | Genetics Research | Cambridge Core
Imputation of single nucleotide polymorphism genotypes in biparental, backcross, and topcross populations with a hidden Markov model

Mosaic analysis of androgenetic/biparental cell lines using STR markers... | Download Scientific Diagram
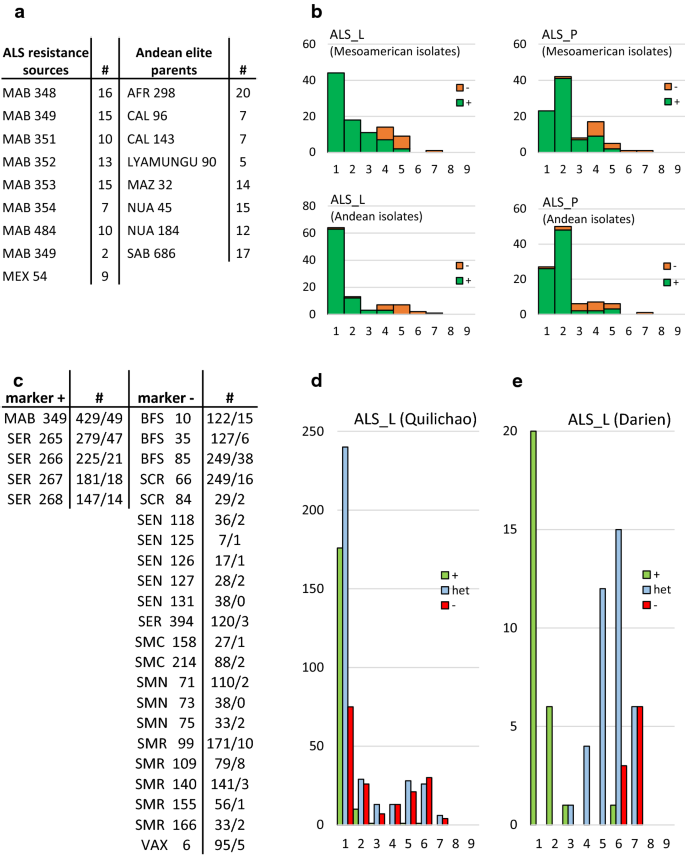
Fine-mapping of angular leaf spot resistance gene Phg-2 in common bean and development of molecular breeding tools | SpringerLink
DNA Marker Transmission and Linkage Analysis in Populations Derived from a Sugarcane (Saccharum spp.) x Erianthus arundinaceus Hybrid | PLOS ONE
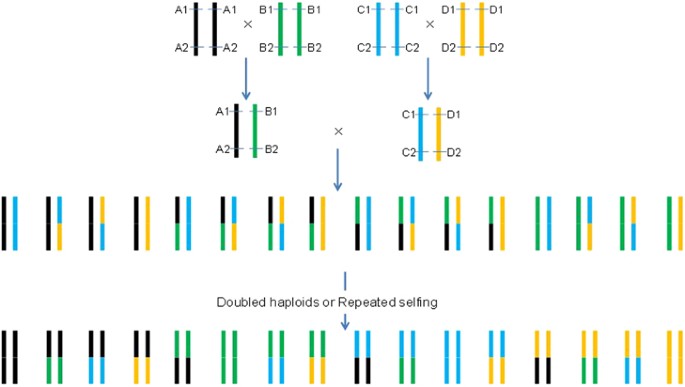
Background controlled QTL mapping in pure-line genetic populations derived from four-way crosses | Heredity
![PDF] Bi-parental cytoplasmic DNA inheritance in Wisteria (Fabaceae): evidence from a natural experiment. | Semantic Scholar PDF] Bi-parental cytoplasmic DNA inheritance in Wisteria (Fabaceae): evidence from a natural experiment. | Semantic Scholar](https://d3i71xaburhd42.cloudfront.net/f85c9d5ad27183dd0a164e3e37b3b05c588a051c/2-Table1-1.png)
PDF] Bi-parental cytoplasmic DNA inheritance in Wisteria (Fabaceae): evidence from a natural experiment. | Semantic Scholar
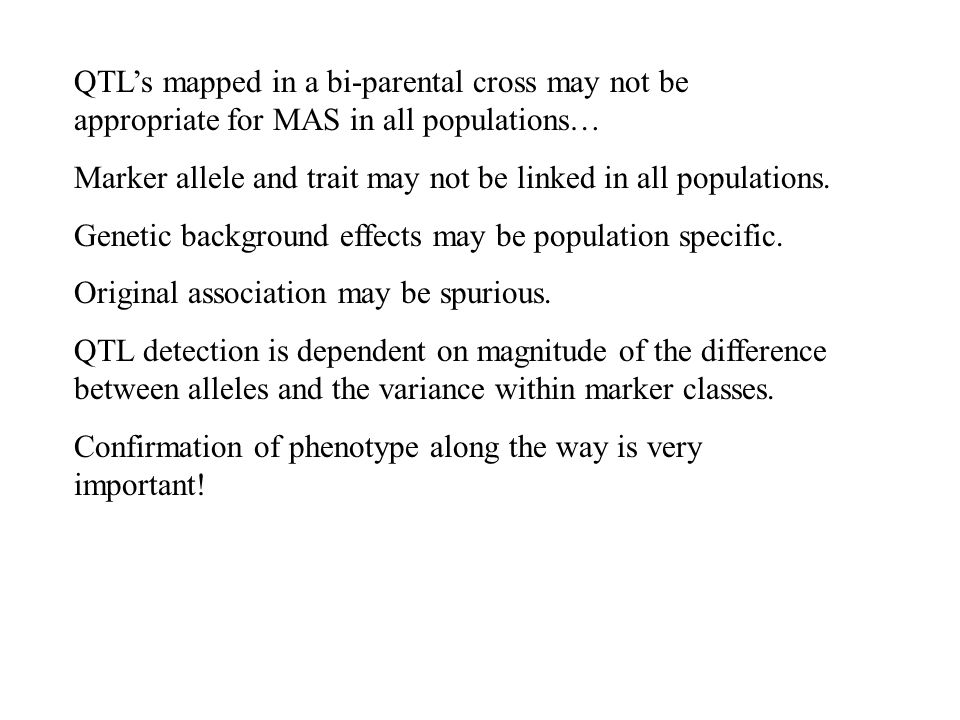

Software and Databases for managing and selecting molecular markers General introduction Pathway approach for candidate gene identification and introduction. - ppt download

Bridging the genotyping gap: using genotyping by sequencing (GBS) to add high-density SNP markers and new value to traditional bi-parental mapping and breeding populations
A novel resistance gene for bacterial blight in rice, Xa43(t) identified by GWAS, confirmed by QTL mapping using a bi-parental population | PLOS ONE

Figure 1 | Biparental Crossing and QTL Mapping for Validation of Genome-Wide Association Studies | SpringerLink

High-density SNP mapping reveals closely linked QTL for resistance to Stagonospora nodorum blotch (SNB) in flag leaf and glume of hexaploid wheat